| Class | Analysis and data manipulation command |
| Name | predict |
| Arguments | <ytrait> on <loc1>...[to]...<locN> [complete_obs]. |
Replace quantitative trait with the predicted value from the multivariate regression on the list of predictors (which may include the average allele length of a marker locus). The com option means only individuals with no missing values for any of the listed traits will be updated. Otherwise, missing values are replaced with the sample mean for that phenotype, when calculating the predicted value.
Example:
>> predict predChol = age ldl
------------------------------------------------
Linear regression analysis of trait "predChol"
------------------------------------------------
Variable Beta Stand Error t-Value
-----------------------------------------------------
Intercept 238.1684 43.0137 5.5370 ***
age -3.8292 1.5887 2.4102 *
ldl*2 120.6935 35.4199 3.4075 **
No. usable observations = 27 ( 45.0%)
Model Mean Square = 74284.9308 (df= 2)
Mean Square Error = 11872.6848 (df= 24)
Multiple R**2 = 0.3427
Akaike Inf. Criterion = 9.5301
Wrote 60 predicted values to predChol.
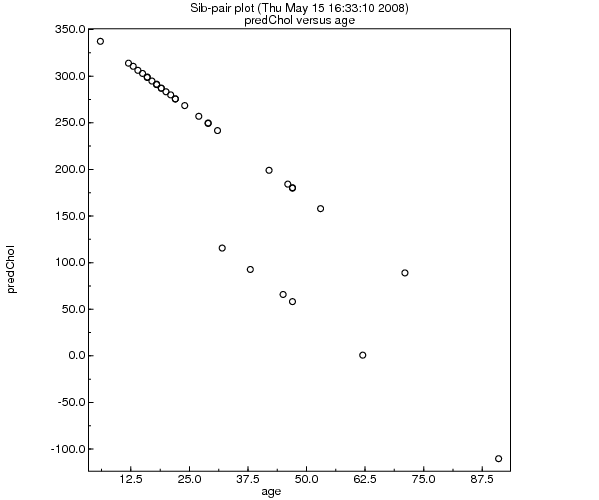
>> plot age predChol

| << (residuals) | Up to index | >> (impute) |