>> include linclex.in
>> set locus pc{1 2} qua
>> mds pc1 pc2
------------------------------------------------
IBS marker sharing: first 2 principal components
------------------------------------------------
Pairwise mean identity-by-state sharing based on 1 to 3 markers.
There are 66 potentially useful individuals.
1.000
0.500 1.000
0.500 0.500 1.000
0.500 0.500 1.000 1.000
0.500 1.000 0.500 0.500 1.000
0.667 0.500 0.167 0.167 0.500 1.000...
Extracting principal components
Writing loadings on first 2 components to: "pc1" "pc2".
>> head
! f m s l o
! a o e i n D13S162 D13S160 D13S170
! id t t x n e pc1 pc2
!
1 1 x x m n n 2/4 1/5 5/9 -0.0256 0.0066
1 2 x x f n n 4/5 3/5 5/7 -0.0277 -0.0882
1 3 1 2 f y y 4/4 5/5 5/5 0.2296 -0.0353
1 4 1 2 m y n 4/4 5/5 5/5 0.2296 -0.0353
1 5 1 2 f n n 4/5 3/5 5/7 -0.0277 -0.0882
1 6 1 2 f n n 2/4 1/3 7/9 -0.1341 0.1207
1 7 1 2 f n n 2/4 1/3 7/9 -0.1341 0.1207
2 1 x x m n n 4/8 2/5 3/5 -0.0793 -0.0490
2 2 x x f n n 4/6 4/5 1/5 -0.0545 -0.1477
2 3 1 2 m n n 6/8 2/4 1/3 -0.2572 -0.0111
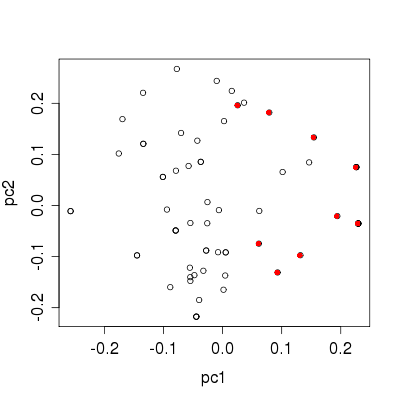
>> plot pc1 pc2
Plot of MDS configuation on first two dimensions. The coloured points represent the cases.